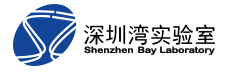Huang, Kai
黄恺
Institute of Systems and Physical Biology
Junior Principal Investigator
huangkai@szbl.ac.cn
Home page of research group:http://huangkai-lab.szbl.ac.cn/
Timeline
-
2020 - Present
Shenzhen Bay Laboratory Junior Principal Investigator
-
2015 - 2020
Northwestern University Postdoc Fellow
-
2009 - 2015
University of Wisconsin-Madison PhD
-
2005 - 2009
Tsinghua University Bachelor
Research Areas
Our group focuses on theoretical computation of biological and biomimetic systems, and the main research directions include, but are not limited to.
(1) Chromatin folding: Combining computer simulations, biomimetic analysis and machine learning methods to predict the advanced structure of chromatin and its regulation of gene expression.
(2) Biological phase separation: theoretical calculations to reveal the physicochemical mechanisms of phase separation and phase transition in biological systems and their correlation with biological functions.
(3) Biomimetic system design: design of novel materials and artificial intelligence algorithms based on the inspiration of biological systems.
Highlights
Proposed a four-dimensional forest model of chromatin folding and a phase-separated higher-order catalytic model of genomic polysome interactions; Revealing the functional structure of disordered regions within the nuclear pore and explaining the pathway selection for trans-nuclear pore transport; Designed and studied multi-functional multi-response intelligent artificial nanopore and high performance ion rectifier; Developed a high-precision and high-efficiency theoretical calculation method for non-equilibrium states. This work has been published in leading international journals such as Science Advances, Nature Communications, Journal of the American Chemical Society, ACS Nano, and Cell Reports. The work on the advanced structure of the genome has received special attention from Francis Collins, Director of the NIH and former head of the Human Genome Project, who commented that "the new picture of the human three-dimensional genome provides new insights for understanding and attacking cancer".
Honors
• 2020 Shenzhen Peacock Project Talent
• 2022 Pearl River Talents Program of Guangdong Province Young Top Talent Project
Related News
1. NIH Director's Blog | A New View of the 3D Genome
2. PHYS ORG | Chromatin organizes itself into 3-D 'forests' in single cells
3. SOTT | Chromatin organizes itself into 3D 'forests' in single cells
4. PHYS ORG | Researchers settle long-standing debate about fundamental behavior of shaking particles
5. IFLScience | Disorder May Be In Your DNA And Affecting Your Chance Of Surviving Cancer
Selected Publications
1. Z. Shi, J. Xu, L. Niu, W Shen, S. Yan, Y. Tan, X. Quan, E. Cheung, K. Huang*, Y. Chen*, L. Li*, C. Hou*. Evolutionarily distinct and sperm-specific supersized chromatin loops are marked by Helitron transposons in Xenopus tropicalis. Cell Reports, 2023, 42(3):112151. (*co-corresponding author)
2. A. R. Shim, K. Huang*, V. Backman, I. Szleifer*. Chromatin as self-returning walks: From population to single cell and back. Biophysical Reports, 2022, 2(1), 100042.(*co-corresponding author)
3. J. Wang, Y. Tao, Y. Juan, H. Zhou, X. Zhao, X. Cheng, X. Wang, X. Quan, J. Li, K. Huang*, W. Wei*, J. Zhao*. Hierarchical Assembly of Flexible Biopolymer Polyphosphate-Manganese into Nanosheets. Small, 2022, 18(41):e2203200.(*co-corresponding author)
4. S. Qin, K. Huang*, I. Szleifer*. Design of Multifunctional Nanopore Using Polyampholyte Brush with Composition Gradient. ACS Nano, 2021, 15(11):17678-17688. (*co-corresponding author)
5. K. Huang*, Y. Li, A. R. Shim, R. K. A. Virk, V. Agrawal, A. Eshein, R. J. Nap, L. M. Almassalha, V. Backman*, I. Szleifer*. Physical and data structure of 3D genome. Science Advances, 2020, 6(2):eaay4055. (*co-corresponding author).
6. K. Huang, M. Tagliazucchi, S.H. Park, Y. Rabin and I. Szleifer, “Nanocompartmentalization of the nuclear pore lumen”, Biophysical Journal, 118, 219-231 (2020)
7. K. Huang, I. Szleifer. Design of Multifunctional Nanogate in Response to Multiple External Stimuli Using Amphiphilic Diblock Copolymer. Journal of the American Chemical Society, 2017, 139(18):6422-6430.
8. K. Huang*, I. Szlufarska. Effect of interfaces on the nearby Brownian motion. Nature Communications, 2015, 6(1):8558.(*co-corresponding author)