汇天下英才 建世界舞台
李刚
Gang LI
系统与物理生物学研究所
Timeline
-
2020-Present
Shenzhen Bay Laboratory Principal Investigator
-
2016-2020
Institute of Genomics and System Biology & Chemistry Department at the University of Chicago Postdoc in Chemical Biology and Cancer Research
-
2010-2016
College of Chemistry and Molecular Engineering, Peking University Ph. D. in Chemistry
Research Areas
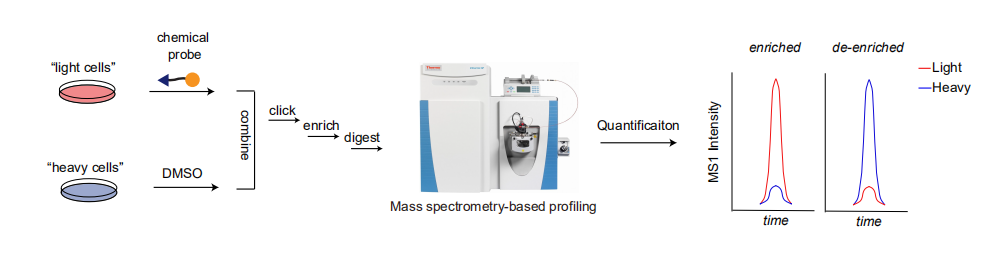
Research in Li lab lies at chemical proteomics and nucleic acid chemical biology, with an eye towards intervening in human diseases: 1) chemical probe design based on reactive metabolites, enzyme cofactors, and bioactive compound for proteomic profiling; 2) system biology platform development for novel biological pathway elucidation; 3) multiplexed activity profiling for biomarker discovery in clinical samples; 4) DNA-encoded small molecule library selection for specific enzyme inhibitor/probe discovery.

Achievements
Gang developed targeted chemical proteomics platforms for spatially resolved enzyme activities mapping in single cells as well as ultrasensitive biomarker activity profiling in clinical samples. He is an inventor on a patent for multiplexed biomarker activity profiling in clinical samples for prognosis. He also built up a novel target identification platform using DNA-templated chemistry and an inhibitor discovery platform by integrating DNA-encoded dynamic library with Illumina sequencing. Gang’s work has been published in high impact journals such as Nature Communications, PNAS, Angewandte Chemie and Chemical Science.
Honors
• National Scholarship (China)
• Principle Scholarship (Peking University)
• Kharasch Travel Award (University
Current Openings
1. 博士后
2. 研究助理
Selected Publications
1. Gang Li, Mark A. Eckert, Jae W. Chang, Jeffrey E. Montgomery, Agnieszka Chryplewicz, Ernest Lengyel, Raymond E. Moellering*, Ultrasensitive, multiplexed chemoproteomic profiling with soluble activity-dependent proximity ligation, Proc. Natl. Acad. Sci. U.S.A. 2019, 116, 21493-21500.
2. Gang Li, Jeffrey E. Montgomery, Mark A. Eckert, Jae W. Chang, Sam M. Tienda, Ernest Lengyel, Raymond E. Moellering* , An activity-dependent proximity ligation platform for spatially resolved quantification of active enzymes in single cells, Nat. Commun. 2017, 8, 1775-1787.
3. Gang Li, Ying Liu, Yu Liu, Li Chen, Siyu Wu, Yang Liu, Xiaoyu Li*, Photoaffinity labeling of small-molecule-binding proteins by DNA-templated chemistry, Angew. Chem. Int. Ed. 2013, 52, 9544-9549.
4. Gang Li, Wenlu Zheng, Zitian Chen, Yu Zhou, Yu Liu, Junrui Yang, Yanyi Huang, Xiaoyu Li* , Design, preparation, and selection of DNA-encoded dynamic libraries, Chem. Sci. 2015, 6, 7097-7104.
5. Gang Li, Wenlu Zheng, Ying Liu, Xiaoyu Li* , Novel encoding methods for DNA-templated chemical libraries, Curr. Opin. Chem. Biol. 2015, 26, 25-33.